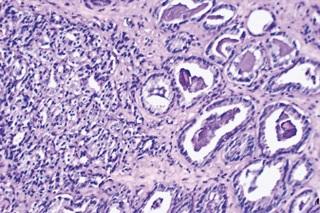
Prostate cancer metastasis linked to revival of dormant molecular program
- Researchers generated the world’s largest epigenomic dataset in prostate biology
- Data confirm decades-old hypothesis that metastatic disease hijacks tissue-specific developmental programs
When prostate cancer progresses to a more-dangerous metastatic state, it does so by resurrecting dormant molecular mechanisms that had guided the fetal development of the prostate gland but had been subsequently switched off, say scientists from Dana-Farber Cancer Institute.
The study, an international collaboration with The Netherlands Cancer Institute, was published in Nature Genetics. “It shows that particular programs that were operative during prostate fetal development become reactivated during metastatic disease,” said Matthew Freedman, MD, a Dana-Farber medical oncologist and co-corresponding author of the report. “The reactivation of these programs is presumably important for the spread of the disease, and if we could understand it better, and potentially block or inhibit the process, it may help us to suppress metastatic prostate disease.”
The insight, gained from one of the largest studies of charting the epigenetic landscape in normal, cancerous, and metastatic prostate tissues, could lay the groundwork for identifying ways of slowing or preventing the initiation and spread of prostate cancer, the investigators say. The comprehensive study of epigenetic changes in prostate cancer, the authors propose, “is foundational for understanding the mechanisms underlying tumor progression” and identifying viable therapeutic targets and vulnerabilities.
The discovery reflects the crafty, efficient nature of cancer’s survival strategies. The investigators showed that cells in a localized prostate tumor need not adopt a new mechanism to acquire the characteristics that enable them to migrate away from their origin and travel around the body. Rather, they reactivate a molecular pathway which, in the developing embryo and fetus, had enabled cells to move about and invade tissues to form the prostate gland in fetal development. Then, no longer needed, the pathway was shut down in the mature prostate gland. Although silenced, the molecular “memory” of the mechanism remained present in the epigenome of the prostate cells.
“As the cancer cell searches for ways to grow and metastasize, it appears to access and hijack the program that is most easily ‘visible’ – the bookmarked developmental site,” explained Mark Pomerantz, MD, co-first author of the report. He is a medical oncologist in the Lank Center for Genitourinary Oncology at Dana-Farber. “Development is a time of great movement and a time of great invasion – similar traits that are used by tumor cells to metastasize,” explained Freedman.
“We were amazed to see this was a universal phenomenon, shared by all metastatic prostate tumors we studied,” said Wilbert Zwart, PhD, of the Netherlands Cancer Institute and co-corresponding author of the study.
Prostate cancer can be viewed as an epigenetic disease. Epigenetics literally means “above” or “on top of” genetics and refers to modifications that turn genes “on” or “off” without changing the DNA sequence.
The investigators say that the study produced the largest epigenomic dataset in prostate biology spanning normal, prostate tumors, as well as metastatic prostate disease. “It’s a landscape that will serve as a foundation for countless other investigators to utilize these data within their own experiments,” said Pomerantz.
The researchers analyzed 268 sets of epigenetic data from patient-derived samples related to the transition of normal prostate tissue to localized prostate tumors, and from localized tumors to metastatic disease. They integrated the datasets to identify thousands of sites across the prostate genome that regulate expression of nearby and faraway genes. The specific reconfiguration of these gene regulatory elements is implicated in prostate tumor formation and progression. To demonstrate the broad utility of the data, the researchers also noted that the DNA code within these regulatory elements, “were strongly associated with prostate cancer risk heritability” – that is, these regions harbored genetic variants that increase the likelihood of someone developing prostate cancer.
“This work has been the result of many years of intense international collaboration and a prime example of how international teamwork pushes the field forward to better understand cancer,” said Zwart.
This study was funded by Rebecca and Nathan Milikowsky, a Prostate Cancer Foundation Challenge Award, the National Institutes of Health (grants R01GM107427, R01CA193910, K0813CA218530), VIDI grant from the Netherlands Organisation for Scientific Research, the Dutch Cancer Society/Alpe d'HuZes (10084) and Oncode Institute, Jean Perkins Foundation, Prostate Cancer Foundation, STOP Cancer Foundation, the U.S. Department of Defense (grant W81XWH-14-1-0273), the H.L. Snyder Foundation, and the National Cancer Institute/National Institutes of Health (grant P50CA092131).
The Pacific Northwest Prostate Cancer SPORE (grant P50 CA097186), the Department of Defense (grant W81XWH-17-1-0415, P01 CA163227), and the Institute for Prostate Cancer Research supported establishment and generation of the LuCaP PDXs models.
Media Contacts
If you are a journalist and have a question about this story, please call 617-632-4090 and ask to speak to a member of the media team, or email media@dfci.harvard.edu.
The Media Team cannot respond to patient inquiries. For more information, please see Contact Us.